Scratchpad
This is where we fiddle around with our data and code.
0.1 Load packages
0.2 Working with open scholarly metadata
# Get list of refs from the OpenCitations API.
# Get list of refs from the OpenCitations API
response <- GET("https://opencitations.net/index/api/v2/references/doi:10.1007/s10814-021-09163-3")
content <- content(response, "text")
parsed_data <- fromJSON(content)
seed_refs <- sub('.*doi:', '', parsed_data$cited)
seed_refs <- gsub( " .*$", "", seed_refs)
seed_refs <- data.frame(seed_refs)
colnames(seed_refs)[1] <- "doi"# Retrieve bibliographic metadata for each referenced work from Crossref.
seed_refs_cr <- cr_works(seed_refs$doi)
seed_refs_cr <- seed_refs_cr$data
seed_refs$title <- seed_refs_cr$title[match(seed_refs$doi, seed_refs_cr$doi)]
seed_refs$container_title <- seed_refs_cr$container.title[match(seed_refs$doi, seed_refs_cr$doi)]
seed_refs$type <- seed_refs_cr$type[match(seed_refs$doi, seed_refs_cr$doi)]
seed_refs$date <- seed_refs_cr$created[match(seed_refs$doi, seed_refs_cr$doi)]
seed_refs$year <- substr(seed_refs$date, 1, 4)
# It's not necessary to retrieve abstracts via API.
# Abstracts are too messy and inconsistent, and it's a pain to deal with html tags.# Generate lists of authors, indexed by DOIs.
# TBDI’m missing a step here, from my previous work several months ago. Need to find the code that generated seed_refs_bib.
# Filter bib file for items with a DOI.
seed_refs_bib <- bib2df::bib2df(paste("https://opencitations.net/index/api/v2/references/doi:",seed_refs_bib[!is.na(seed_refs_bib$DOI),]$DOI))# Query CrossRef for metadata pertaining to references with a DOI.
f1_cr <- cr_works(seed_refs_bib[!is.na(seed_refs_bib$DOI),]$DOI)
f1_cr <- f1_cr$data# Query OpenCitations for references pertaining to f1.
f1_oc <- oc_coci_meta(seed_refs_bib[!is.na(seed_refs_bib$DOI),]$DOI)
response <- GET("seed_refs_bib[!is.na(seed_refs_bib$DOI),]$DOI")
content <- content(response, "text")
parsed_data <- fromJSON(content)
f1_oc <- sub('.*doi:', '', parsed_data$cited)
f1_oc <- gsub( " .*$", "", f1_oc)
f1_oc <- data.frame(f1_oc)
colnames(f1_oc)[1] <- "doi"# For data cleaning purposes.
# Modify the variables to find articles without DOI, non-articles with DOI, etc.
seed_refs_bib %>%
filter(CATEGORY != "ARTICLE",
!is.na(DOI)
)
seed_refs_bib %>%
filter(CATEGORY == "BOOK"
)0.3 Cleaning and integrating the annotations spreadsheet
1 Basic descriptive statistics
Set up some initial color palettes based on the nord package.
Read in the data.
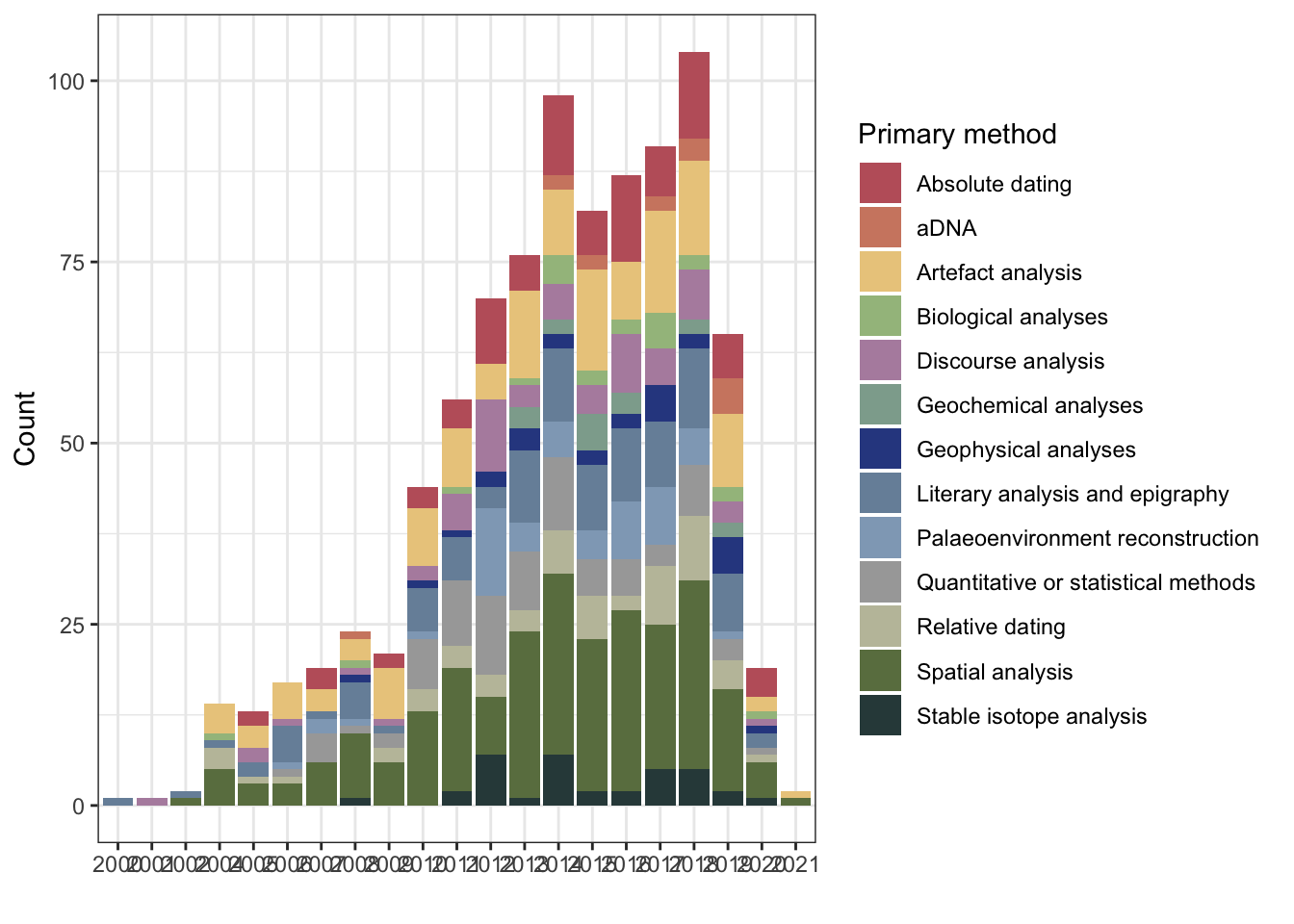
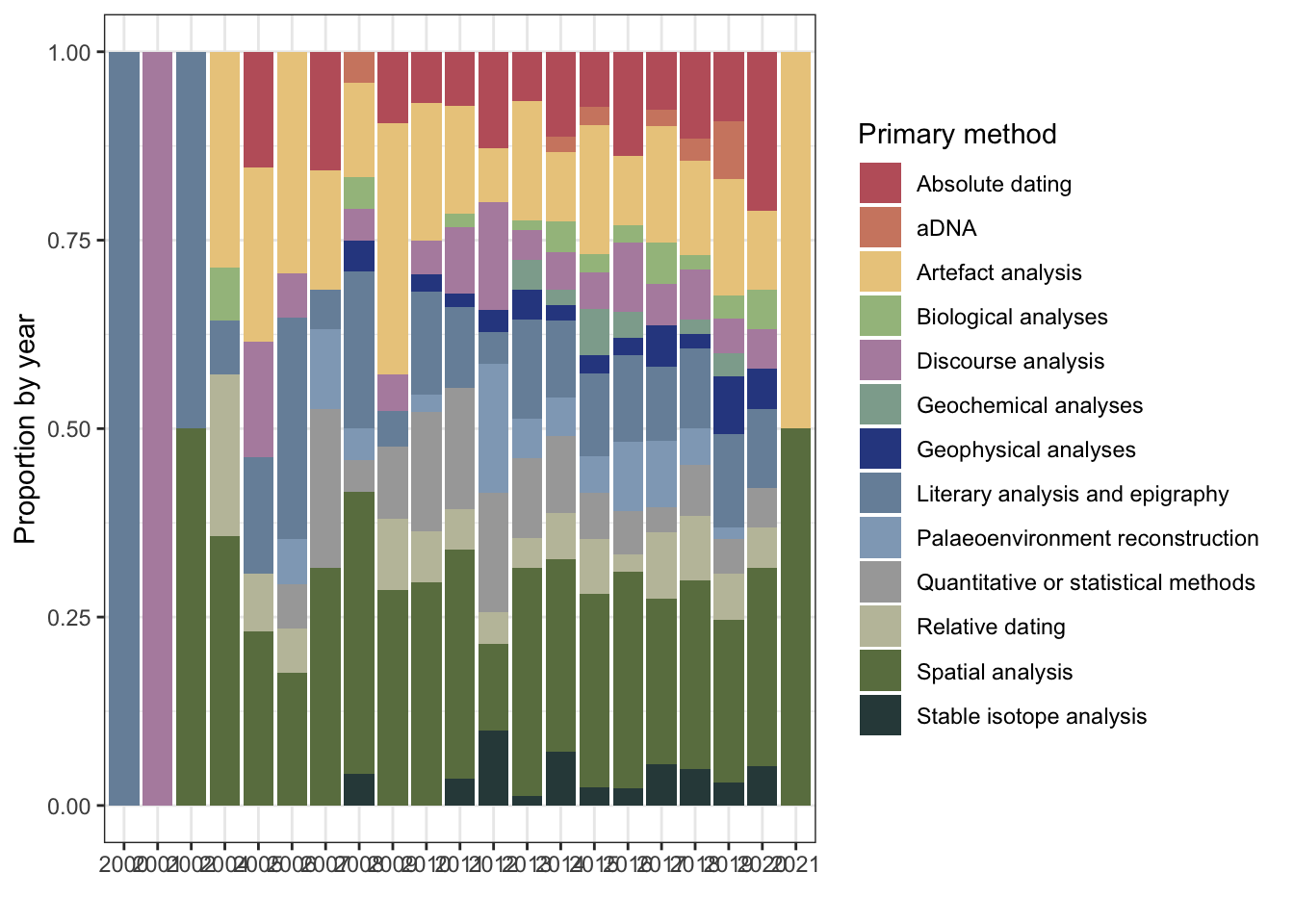
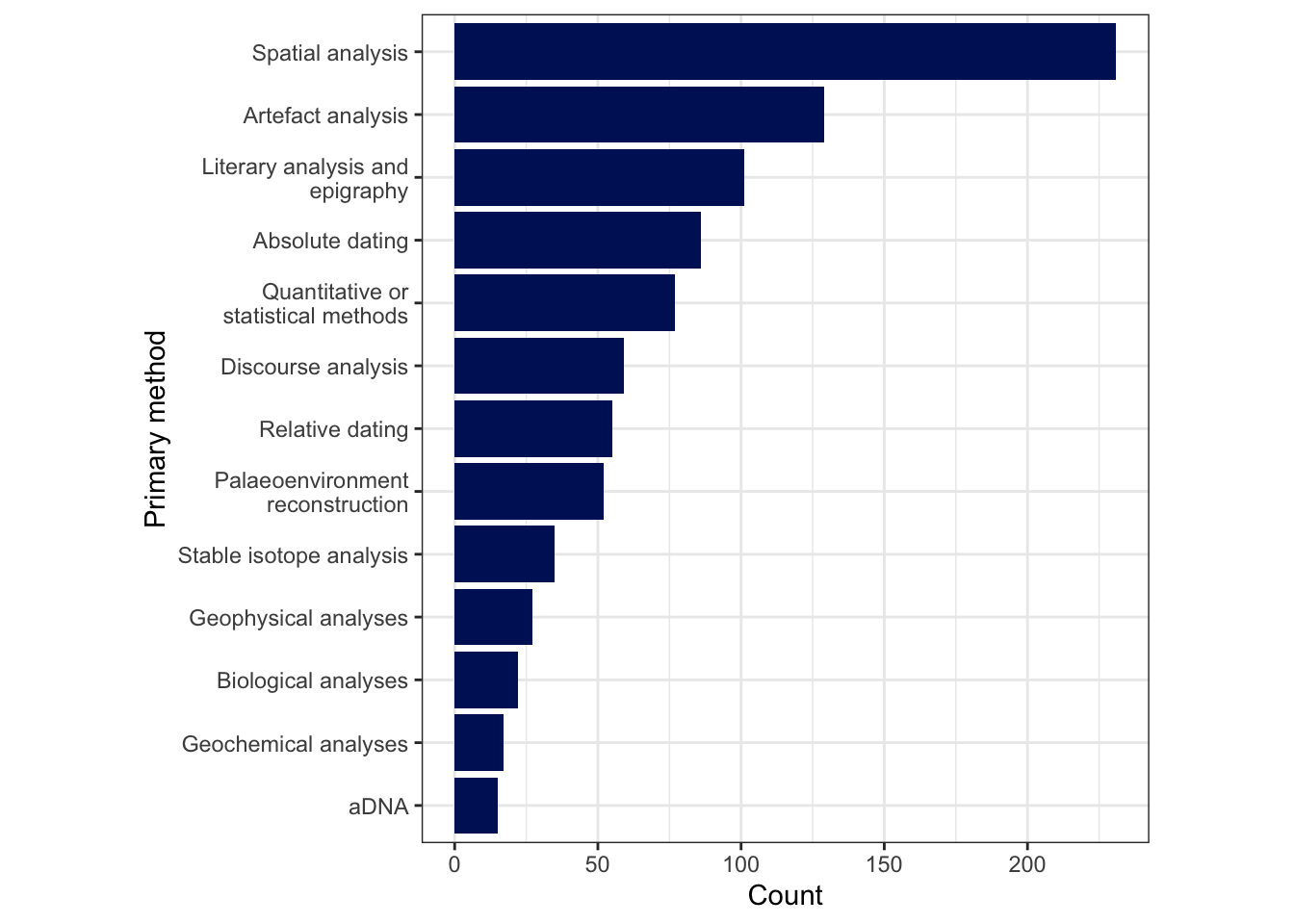
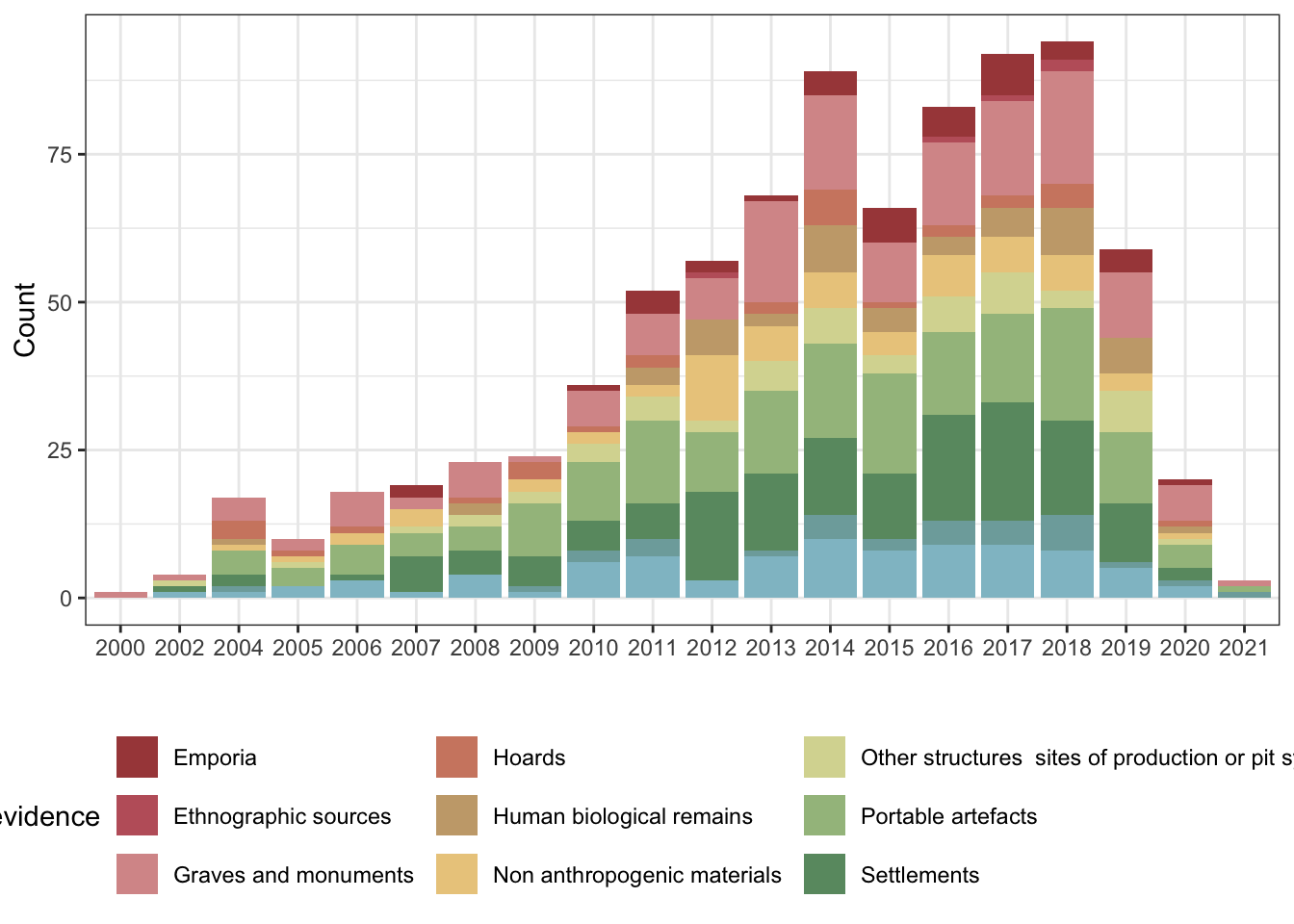
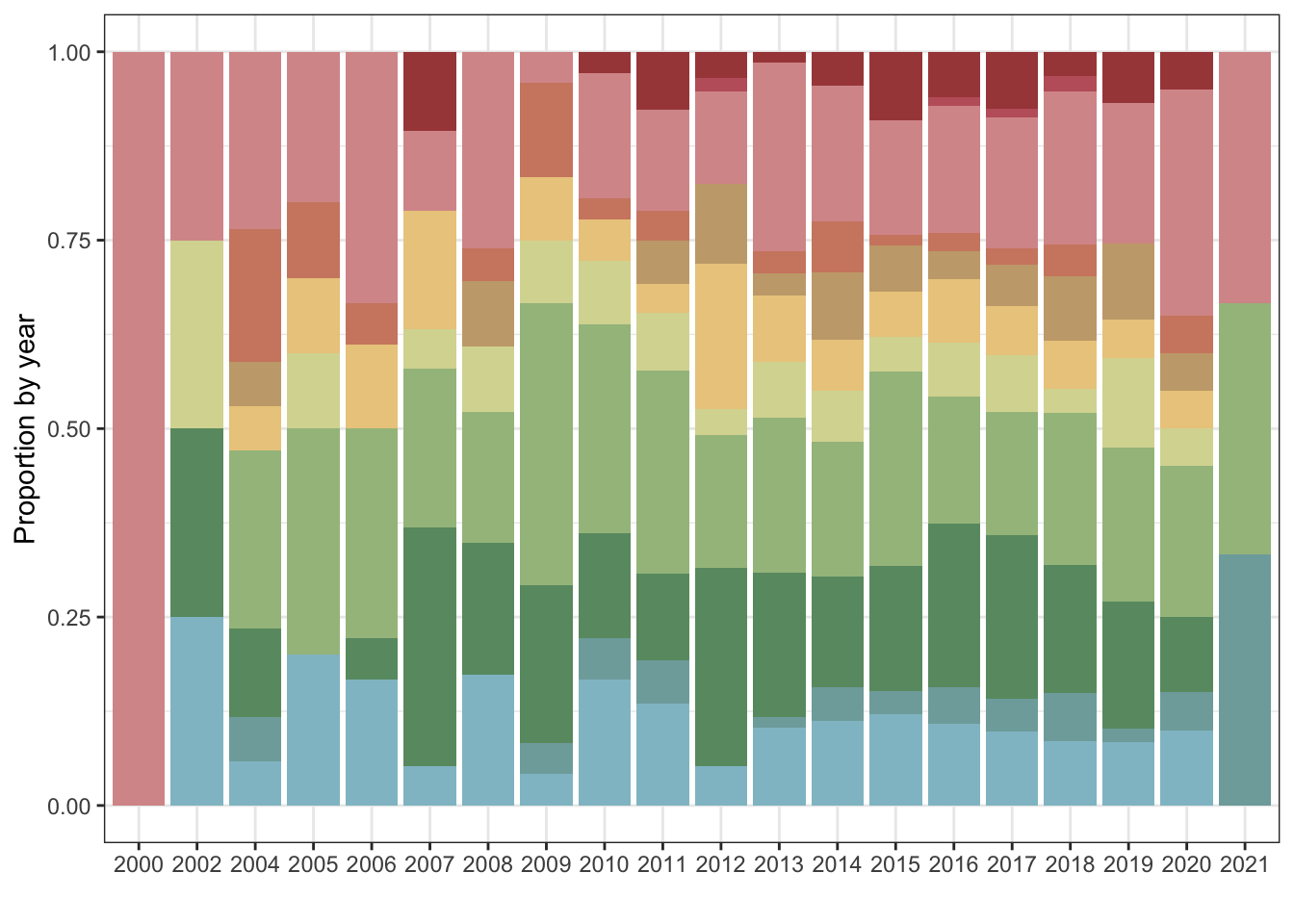
1.1 Intial plots
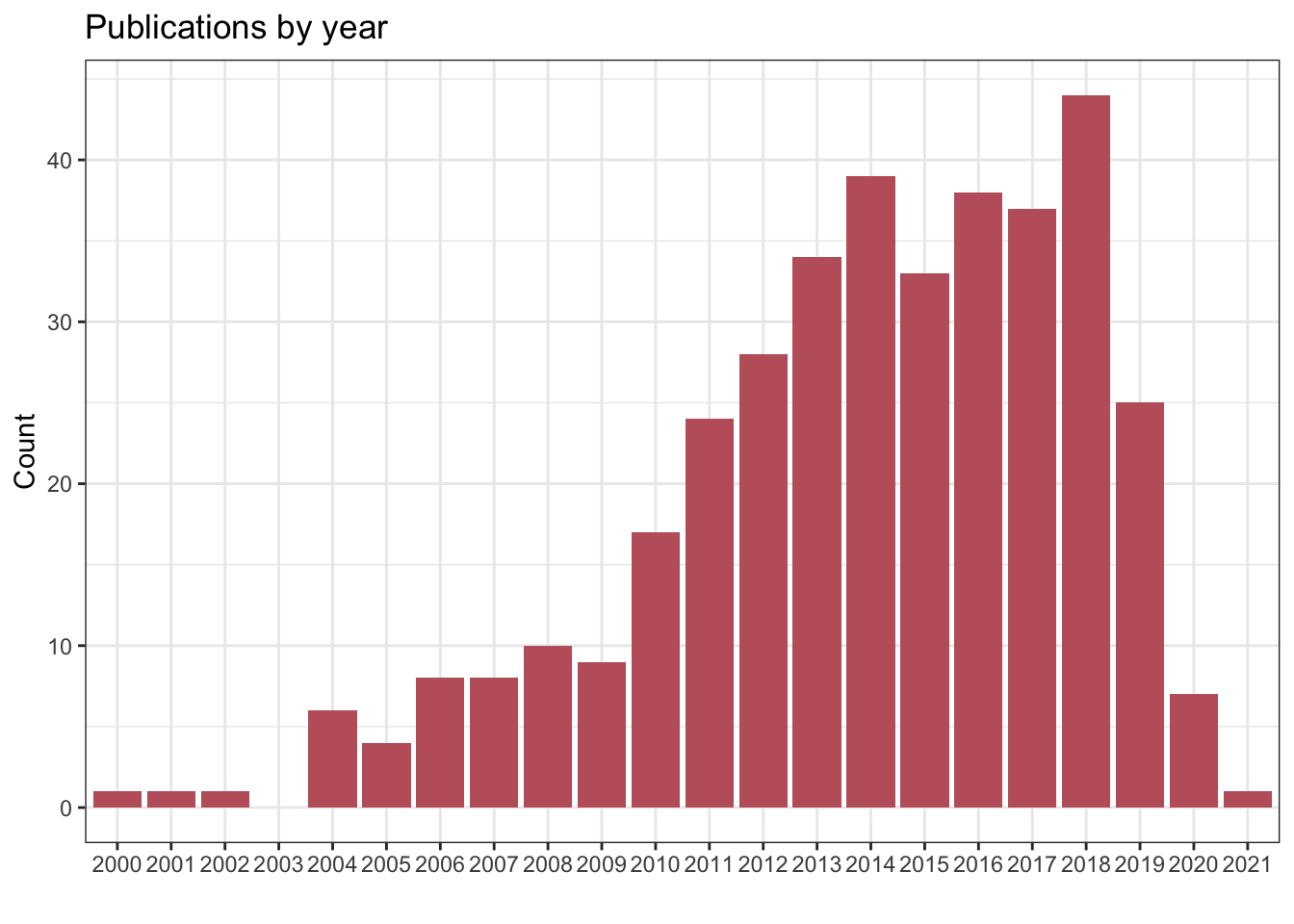
1.1.1 Number of publications per year
1.5 Citation networks
TBD: Integrate the BibVik-CitationAnalysis toolkit here, somehow.
1.6 Citation contexts
TBD: Integrate the BibVik-CitationAnalysis toolkit here, somehow.
1.7 Citation clusters
TBD: Integrate the BibVik-CitationAnalysis toolkit here, somehow.